General options
for processing functions:
For most processing functions, you will get a specific window
for setting parameters and options. Some of those options are
identical for most of the functions:
- At the top, there is a dropdown selector containing the current
spectra list. Here you can select the spectrum to be processed
(if only one spectrum gets treated). If a preview is available,
the spectrum is selected here.
- if the "treat all spectra" option is checked,
all spectra will get processed and attached as new spectra at
the end of the spectrum list. Otherwise, only the currently selected
spectrum gets processed.
- With the "remove all" option checked, the original
spectra will be removed and only the processed spectra/ spectrum
remain(s) .
- The "keep legend" option leaves the original
legend text unchanged. If unchecked, a function-specific text
part will be added to the legend text.
|
|
|
Simple
Baseline subtraction
For offset-correcting
of straight baselines that deviate from zero. After clicking the
button, use the left mouse button in the plot to select the x
axis range which should be set to zero. The mean value of y values
in the selected region will be subtracted from the spectrum as
a constant value. It is done for all spectra at once.
Advanced Baseline
subtraction
For cases, where
a simple offset baseline correction doesn't work, this menu function
gives advanced possibilities with three different baseline creation
algorithms available. Select the baseline algorithm and its parameters
from the button's dropdown menu entry "Baseline Options".
A baseline preview will be shown and updated for the selected
spectrum, based on current settings.
 |
 . .
|
 |
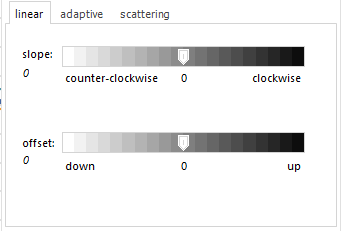
For the
linear baseline, you can adapt these parameters:
- slope: changes the angle of the straight baseline (0 is
horizontal).
- offset: creates a vertical shift. |
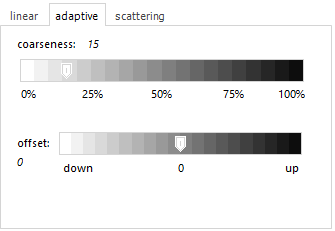
The adaptive
algorithm creates a baseline that snugly fits to the bottom
of spectra. The intention is to remove broad underlying features
(like fluorescence background in Raman spectra), while keeping
actual peaks. A smaller coarseness value creates a tighter
fit, with Offset you can have a vertical shifting of the baseline.
This algorithm is a single-run, no-fitting, non-iterating
algorithm. It creates a baseline by applying a 0% percentile
smoothing, followed by a moving-average smoothing, with the
same interval size for both steps (coarseness translates to
interval size). |
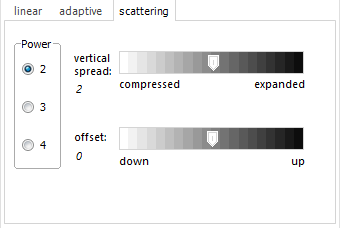
The "scattering"
algorithm is intended for absorbance spectra of scattering
solutions, which have a continuously falling baseline towards
long wavelengths. Depending on the size of scattering centers/
particles, the scattering intensity corresponds to 2nd to
4th power of the wavelength. Additionally, the vertical spread
and offset can be adapted. |
On clicking
"Apply", the created baselines will be subtracted from
the original spectra. With "individual baseline" selected,
a separate baseline is created for each individual spectrum (the
default behaviour). In case of "create baseline spectrum"
is checked, the used baselines will be added as actual spectra
to the list of spectra.
Clicking the main button will execute baseline subtraction on
all spectra, based on the currently defined parameters (but only
if "treat all spectra" is selected).
|
| <jump
back to top> |
|
|
x Offset/ Stretch
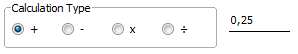
This applies
a constant value with the selected calculation type to the spectrum's
x axis values. Depending on the selected calculation type, you
can shift spectra to left/right (addition (+), subtraction (-)),
stretch (multiplication (x)) and compress (division (÷))
along the x axis.
Hint: Please use this function only if you really know
what you are doing, since it modifies the spectra in a rather
unusual manner.
y Offset/ Stretch
This applies
a constant value with the selected calculation type to the spectrum's
y values. Depending on the selected calculation type, you can
shift spectra up (addition (+)) and down (subtraction (-)), stretch
(multiplication (x)) and compress (division (÷)) along
the y axis. The latter two functions are often used for normalizing
spectra manually.
As an alternative,
you can do both, x and y offsetting and stretching interactively
to individual spectra by using the Scale/Shift
function from the Plot/Views ribbon.
|
| <jump
back to top> |
|

|
Normalization
With normalization,
you bring the y scale of spectra into a similar range, for comparing
spectrum shapes, original y value relations between spectra get
lost. Several methods are available:
Normalize (Peak)
All Spectra
are scaled such that their highest peak in the currently visible
x axis range is set to a value of 1. Select the desired x range
before using the function. It always applies to all loaded spectra
and acts on the spectra, no new spectra will be created.
Normalize (Area)
All Spectra
are scaled such that the area under curve within the selected
x axis range has the same value. After clicking the button, it
waits for you selecting the desired x range with the left mouse
button. It always applies to all loaded spectra and acts on the
current spectra, no new spectra will be created.
Normalize (Value)
This function
will scale spectra, so that they have all a certain value at a
certain x position. Here all the usual options are available,
like treat one or all spectra, remove original, keep legend.
|
| <jump
back to top> |
|
|
Simple
Smooth
For quickly
smoothing all spectra at once, no options available, smoothed
spectra added to the end of spectra list. As smoothing algorithm,
a 3rd order Savitsky-Golay smoothing is used, with an interval
size depending on each spectrum's actual data point step width.
Advanced Smoothing
For better control
of smoothing behaviour, with a choice of smoothing algorithm and
smoothing parameters. Four algorithms are currently available.
Select the smoothing algorithm and its parameters from the button's
dropdown menu entry "Smoothing Options". A preview of
the smoothed spectrum will be shown and updated for the selected
spectrum, based on current settings.
|
|
moving
average smoothes by calculating an average value for
every data point from all neighboring data points within
the selected interval size. A rectangular or triangular
shape can be choosen as weighting function for averaging.
With the triangular shape, more distant data points are
down-weighted.
|
|
|
Savitsky-Golay
smoothes by calculating a least-squares polynomial interpolation
value for every data point from all neighboring data points
within the selected interval size. The interval size and
the polynomial order can be choosen.
|
|
|
Percentile
filter smoothes by identifying the selected percentile
for the neihgbouring data points within the interval size
for every data point. A lower or upper envelope spectrum
results on choosing the 0% or 100% percentile.
|
|
|
Baseline-selective
smoothing is a special algorithm working best on spectra
with baseline areas near to zero, like Raman spectra after
baseline subtraction and some UV/VIS and FTIR spectra. It
is based on Savitsky-Golay smoothing with increasing smoothing
interval size towards the baseline area at low y values.
The gradient of this increase can be changed with the slider
and the mapped y axis range can be defined in two ways.
Just give it a try on your spectra...
The benefit is: have less noise in baseline area without
significantly changing actual peaks!
|
On clicking
"Apply", the defined smoothing algorithm will be applied
on spectra..
Clicking the main button will execute advanced smoothing on all
spectra, based on the currently defined parameters (but only if
"treat all spectra" is selected).
|
| <jump
back to top> |
|
|
Remove
Spikes
With Silicon-based
CCD detectors, artifacts from cosmic radiation are increasing
with longer exposure times (several seconds). These are called
cosmic spikes.
This function
detects spikes and interpolates the spectrum in the range of the
spike. All other spectral data points remain totally unchanged.
Spectra without spikes remain also unchanged. As an option, you
can define the maximum width for a found spike location (default
is 5 pixels). Any found, wider spike artifact is not removed,
to make sure not to change spectral features. Use a smaller value
for spectra with poor resolution, in order to prevent erroneous
removal of narrow spectral peaks. Another options is "backward
removal" which applies the same algorithm from right to left,
thus occasionally finding some undetected spikes from a first
run.
Hint:
you will find a myriad of cosmic spike removal algorithms in literature,
many of which are rather complicated. I devised this simple algorithm
for my own use around year 2001-2002, it was already part of the
previous Spekwin32 software for many years. The algorithm is based
on the very different shape of a spike artifact compared to a
spectral peak in the spectrum's 1st derivative. Based on this
differentiation, the assigned spike area gets interpolated by
a moving average. For a long time, I couldn't find something comparable
in literature. It seems, this finally
changed in 2018...
Remove a peak
This function
exactly does what its name says. Just enter the peak location
and its FWHM and the spectrum gets interpolated in that area.
Hint:
There are only few uses, where making a measured peak go away
is considered a proper task. Handle with care!
Spectrum Part
Cut off
For cutting
away a part of the spectrum from one or both sides (start and
end). Enter the new start and end values for the remaining range
manually, or else interactively move the two vertical cursors
shown together with the function's window. After applying the
new boundaries with Apply, the window will stay open to
allow more changes. When finished, click the Close button.
|
| <jump
back to top> |
|
|
Correct
System Sensitivity
For correcting
the wavelength-dependent system sensitivity (mainly introduced
by fibers, gratings and detectors) of emission spectrometers (fluorescence,
Raman, radiometry...). Sensitivity correction yields "true
spectra" as output.
Load the correction
curve with the Load correction Curve button (any file format
accepted, data should consist of sensitivity against wavelength).
Spectral data will be divided by the loaded correction curve,
which persists for the current session.
|
| <jump
back to top> |
|
|
Chemometric
preprocessing
For preparing
spectra for chemometric analysis, a number of processing steps
are often used, like file conversion, baseline removal, normalization,
smoothing, derivatives and so on. These three functions are often
used as a data pretreatment especially on NIR spectra before actual
chemometric analysis, in order to remove time-dependent drift
that is irrelevant to actual sample characteristics, but would
interfere with analysis. Recommended reads for in-depth explanation
of such algorithms can be found here,
here
and here.
SNV (standard
normal variates) is often used on spectra where baseline and
pathlength changes cause differences between otherwise identical
spectra. It works by normalizing the averages of whole spectra.
MSC
(multiplicative scatter correction)
has proven to be effective in minimizing baseline offsets and
multiplicative effect. It is achieved by regressing a measured
spectrum against a reference spectrum and then correcting the
measured spectrum using the slope and intercept of this linear
fit. You can either load a reference spectrum or use an average
of the whole data as reference.
Detrending
is performed through subtraction of polynomial fit of baseline
from the original spectrum to remove tilted baseline variation,
usually found in NIR reflectance spectra of powdered samples.
The order of the used polynomial can be selected.
|
| <jump
back to top> |
|
![back to SpectraGryph main page [SpectraGryph]](gryphon_white_green_96.png)
![back to SpectraGryph main page [SpectraGryph]](gryphon_white_green_96.png)